Precise determination of protein extinction coefficients under native and denaturing conditions using SV-AUC | SpringerLink

Impact of external amino acids on fluorescent protein chromophore biosynthesis revealed by molecular dynamics and mutagenesis studies - ScienceDirect
![PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/af80a5d2beee749cff2f5d98dc4a8f0dfdd48074/4-Table3-1.png)
PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar
Engrailed and Hox homeodomain proteins contain a related Pbx interaction motif that recognizes a common structure present in Pbx. - Abstract - Europe PMC

A method to identify and quantify the complete peptide composition in protein hydrolysates - ScienceDirect

Molar absorption coefficients for Trp, Tyr, and cystine based on an... | Download Scientific Diagram

Photometric Quantification of Proteins in Aqueous Solutions via UV-Vis Spectroscopy - Eppendorf Handling Solutions
![PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/af80a5d2beee749cff2f5d98dc4a8f0dfdd48074/3-Table2-1.png)
PDF] Calculation of protein extinction coefficients from amino acid sequence data. | Semantic Scholar
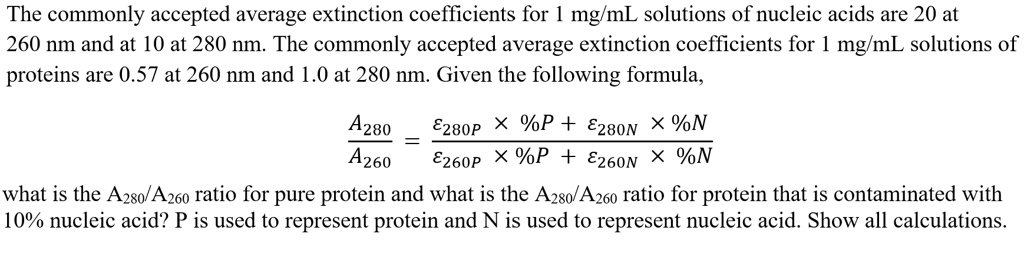

SOLVED: The commonly accepted average extinction coefficients for mg/mL solutions of nucleic acids are 20 at 260 nm and at 10 at 280 nm The commonly accepted average extinction coefficients for mglmL
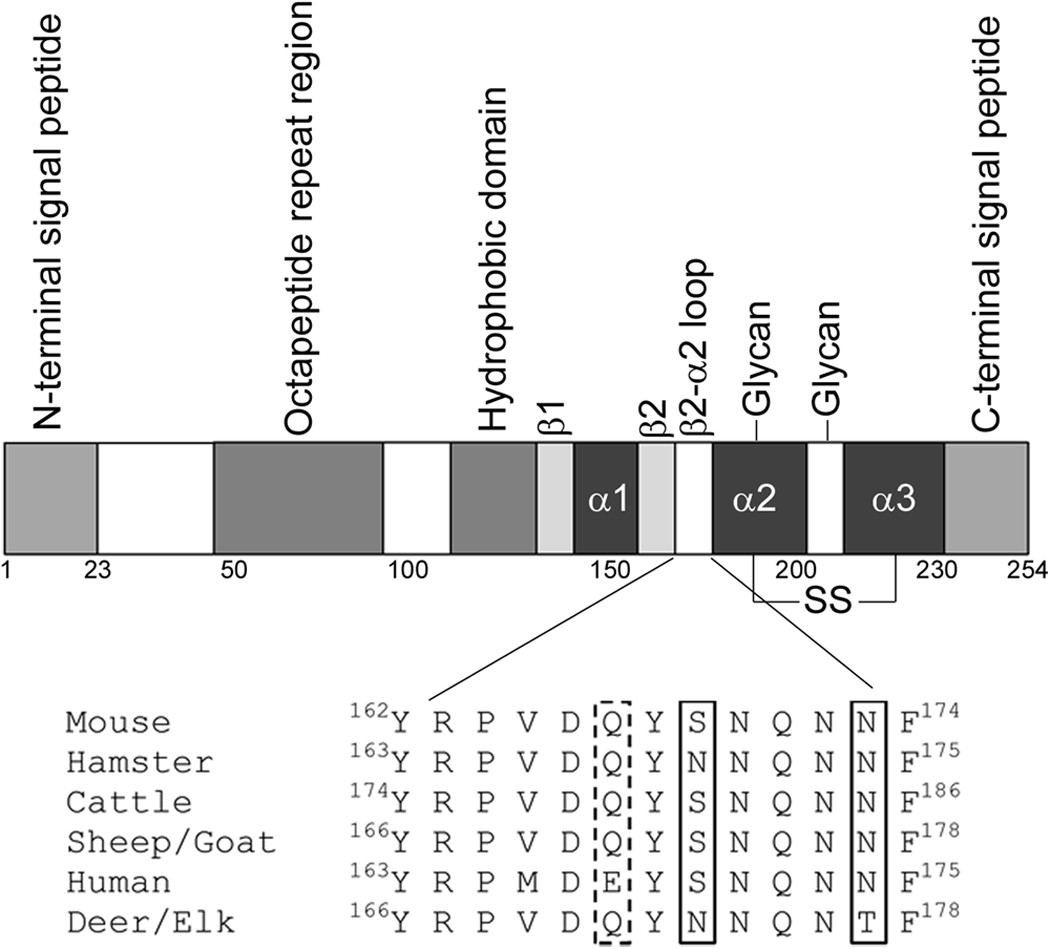
Complex folding and misfolding effects of deer-specific amino acid substitutions in the β2-α2 loop of murine prion protein | Scientific Reports

Molecules | Free Full-Text | Evaluation of Peptide/Protein Self-Assembly and Aggregation by Spectroscopic Methods
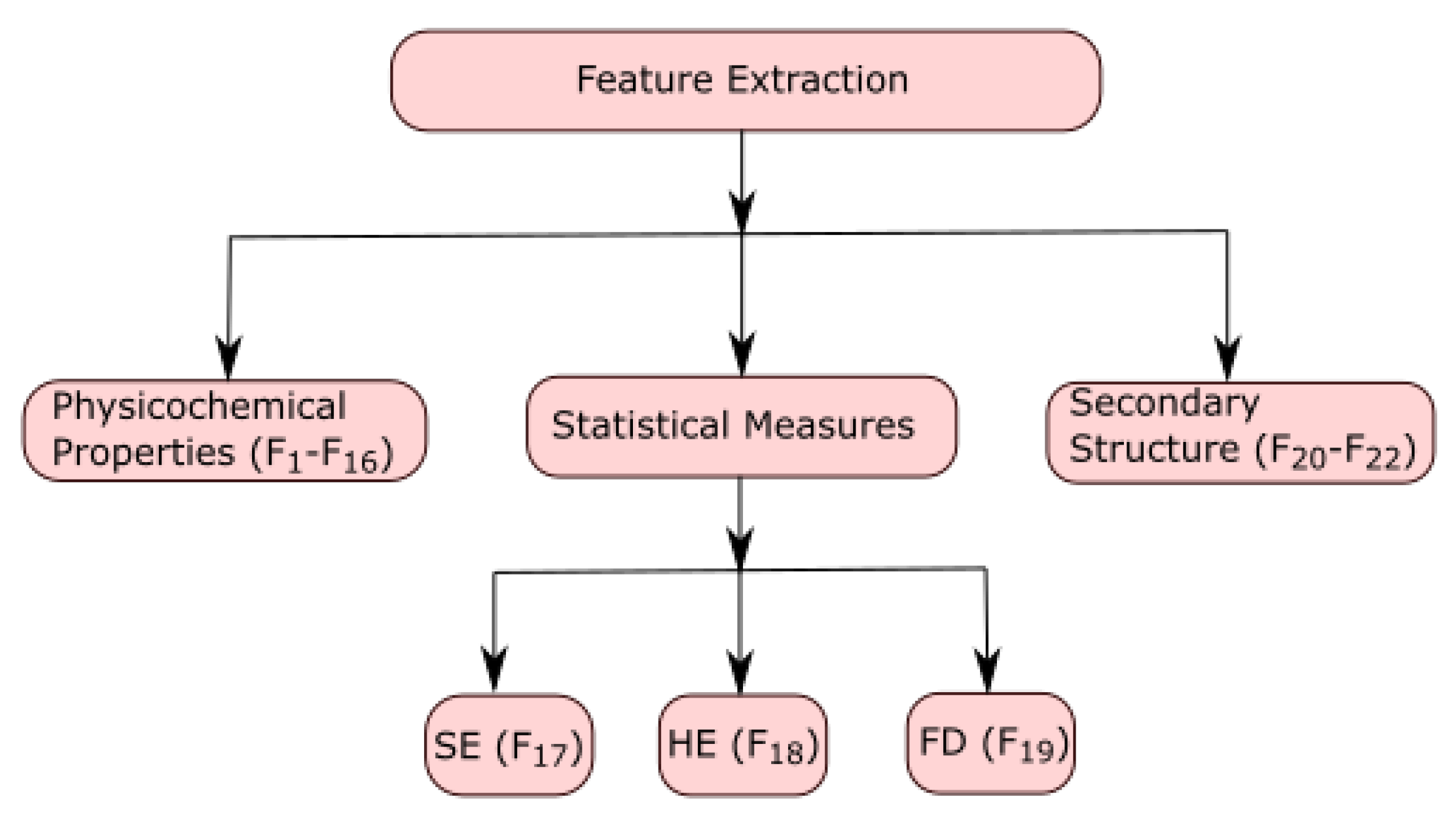
Mathematics | Free Full-Text | Unsupervised Learning for Feature Representation Using Spatial Distribution of Amino Acids in Aldehyde Dehydrogenase (ALDH2) Protein Sequences

The Photophysics of Green Fluorescent Protein: Influence of the Key Amino Acids at Positions 65, 203, and 222: Biophysical Journal